 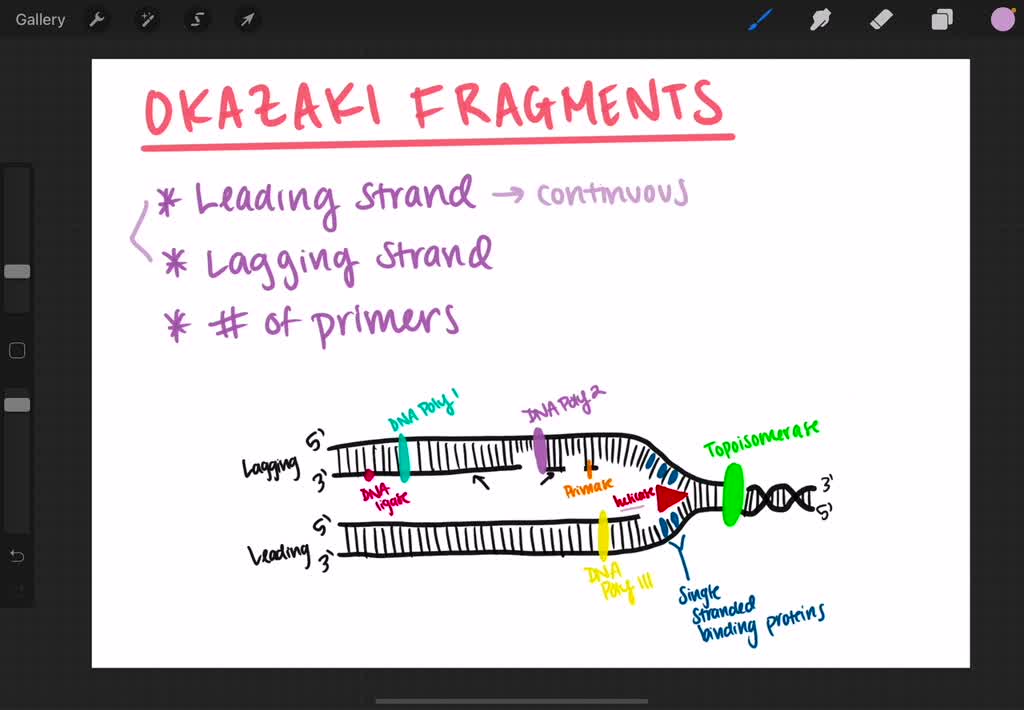
In Escherichia coli, for example, DnaG primase ignores ~97% of the trinucleotide recognition sites and initiates Okazaki fragments only every 1.5–2 kb (and not more frequently) 7. The reason for this flexibility is, however, not understood and is the enigma that we address in the current study: Although extensive research has been carried out on the interactions of DNA primase with DNA, it is still not clear why DNA primase ignores the majority of trinucleotide recognition sites. Importantly, it is now known that even though DNA primase recognizes a specific trinucleotide sequence, flexibility in the selection of initiation sites for Okazaki fragments is allowed 6, i.e., not every primase-DNA recognition sequence (PDRS) will become an Okazaki fragment start site. For example, the activity of bacteriophage T7 DNA primase, comprising the N-domain of gene 4 protein, is initiated by sequence-specific binding of DNA primase to 5’-GTC-3’ 3, 4, which is then followed by the synthesis of a functional primer 5. In prokaryotes, RNA primer formation occurs on pre-defined sequences on the genome that are specifically recognized by a DnaG-type primase 2. This recognition is thus fundamental to the establishment of Okazaki fragments and consequently to the whole process of proper DNA replication. This process of RNA-primed DNA synthesis by a DNA polymerase is triggered exclusively by the recognition of a specific DNA sequence by the primase. On the lagging DNA strand, an enzyme called DNA primase, recognizes the DNA sequence used as the template for the synthesis of RNA primers, and DNA polymerase is then responsible for elongating these RNA primers into the DNA segments known as Okazaki fragments. On the basis of the principles learned about DNA-primase binding, we generated novel DNA sequences with improved binding of T7 primase and improved RNA primer synthesis, as validated experimentally.ĭuring DNA replication the double-stranded DNA is unwound, to expose the two individual DNA strands, one is copied continuously (leading strand) and the other is copied discontinuously (lagging strand). A PBM data learning algorithm enabled the prediction of binding values of T7 primase for any given DNA sequence with unprecedented accuracy and flexibility. To this end, we applied machine-learning algorithms to analyze a massive amount of data obtained from protein-DNA binding microarrays (PBM) with the aim of identifying the essential elements on DNA that are needed for the binding of bacteriophage T7 primase. The unresolved enigma that we address in this study is therefore why some, but not all, primase-DNA recognition sequences (PDRSs) become Okazaki fragment start sites. It has long been known that the frequency of the occurrence of primase trinucleotide recognition on a genome sequence has no influence on the size of the Okazaki fragments. It is therefore reasonable to assume that the binding of DNA primase on a genome marks the start sites of the Okazaki fragments. These RNA primers are elongated by DNA polymerase to form Okazaki fragments on the lagging DNA strand. However, Dna2p has a role in a pathway for processing structured flaps, in which it aids FEN1 using both its nuclease and helicase activities.DNA primase is an essential enzyme that synthesizes short RNA primers on specific DNA sequences. Our results suggest Dna2p is not used for processing of most flaps. 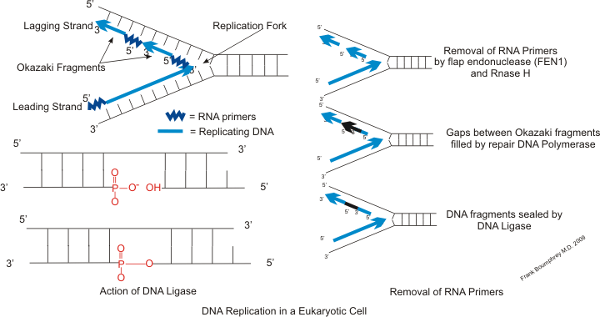
The presence of high Dna2p activity, under reaction conditions favoring helicase activity, substantially stimulated FEN1 cleavage of tailed-foldback flaps and also 30-nucleotide unstructured flaps. CTG flaps can form foldback structures and were inhibitory to both nucleases, however, addition of a dT(12) to the 5'-end of a CTG flap allowed Dna2p cleavage. On 30-nucleotide fixed or equilibrating flaps, RPA partially inhibits FEN1. FEN1 cleaves 10-nucleotide fixed or equilibrating flaps in an efficient reaction, insensitive to even high levels of RPA or Dna2p. To determine the most likely process, we analyzed cleavage of short and long 5'-flaps. RPA dissociates from the resultant short flap, allowing FEN1 cleavage. In the other, the single-stranded binding protein, replication protein A (RPA), coats the flap, inhibits FEN1, but stimulates cleavage by the Dna2p helicase/nuclease. In one model for this process, the flap endonuclease 1 (FEN1) removes the iRNA. Each contains an initiator RNA/DNA primer (iRNA/DNA), which is converted into a 5'-flap and then removed prior to fragment joining. Short DNA segments designated Okazaki fragments are intermediates in eukaryotic DNA replication. On the Roles of Saccharomyces cerevisiae Dna2p and Flap Endonuclease 1 in Okazaki Fragment Processing* Abstract 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
